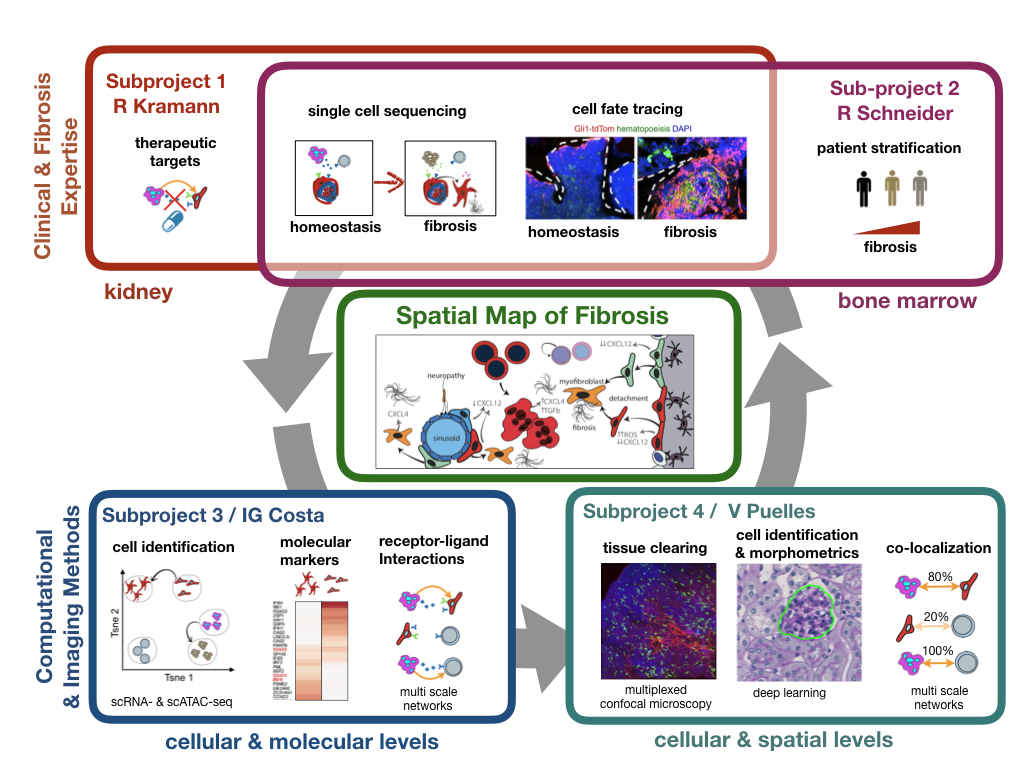
Commentary on the value of multimodal single cell sequencing for inference of gene regulatory networks out in Nature Methods
A short take on how computational approaches leverage multimodal single cell (RNA+ATAC) for inference of regulatory networks and a brief review of recent ground-breaking work by Dictys, SCENIC+ and MIRA at https://rdcu.be/disS3